Creates a choropleth map from an `sf` object, for example one
produced by summarise_points_by_polygon(). Polygons are shaded
according to values in a specified column, with clustering based on the
Fisher–Jenks algorithm.
Usage
choropleth(
data,
value = "output",
id = NULL,
mode = c("plot", "view"),
n = 7,
legend_title = "Value",
palette = "viridis",
id_name = NULL,
...
)Arguments
- data
An object of class
sf.- value
A string giving the name of the column used to shade the polygons.
- id
Optional string giving the name of the column containing polygon IDs used for tooltips.
- mode
A string indicating whether to create a static map (
"plot", default) or an interactive map ("view").- n
Integer; number of clusters. Default is
7.- legend_title
A string giving the legend title.
- palette
A palette name or vector of colors. See
tmaptools::palette_explorer()for available palettes. Prefix the name with"-"to reverse the order. Default is"viridis".- id_name
Deprecated. Use
idinstead.- ...
Additional arguments passed to
tmap::tm_polygons().
Details
The function uses the Fisher–Jenks algorithm
(style = "fisher") to classify values into n groups.
Examples
test <- summarise_points_by_polygon(nl_provincie, insurance, "amount")
#> 109 points are outside any polygon.
choropleth(test, value = "amount_sum")
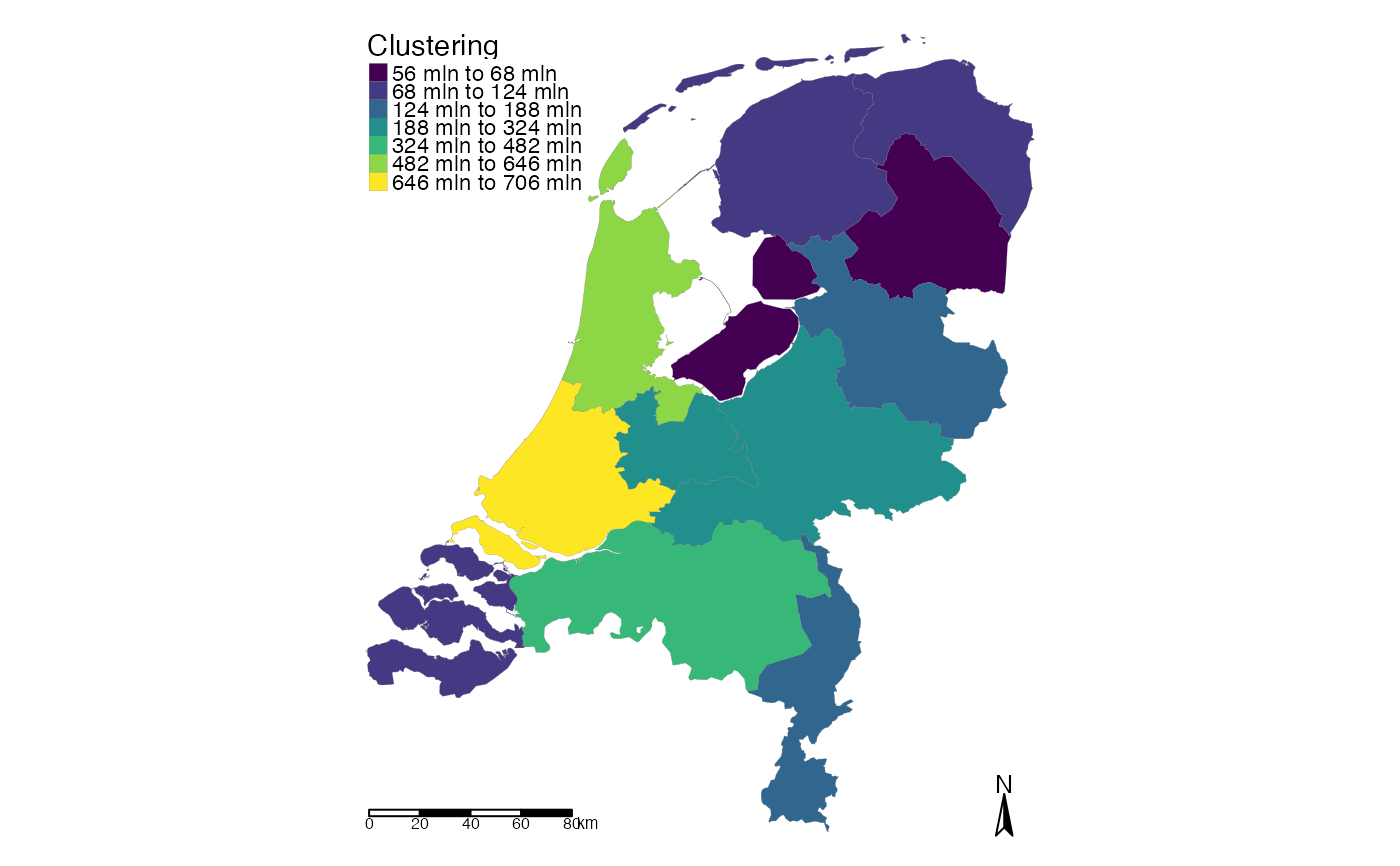 choropleth(test, value = "amount_sum", id = "areaname", mode = "view")
choropleth(test, value = "amount_sum", id = "areaname", mode = "view")
